Technology
The APTASHAPE method is a novel approach for profiling proteins in plasma using RNA aptamers combined with next-generation sequencing.
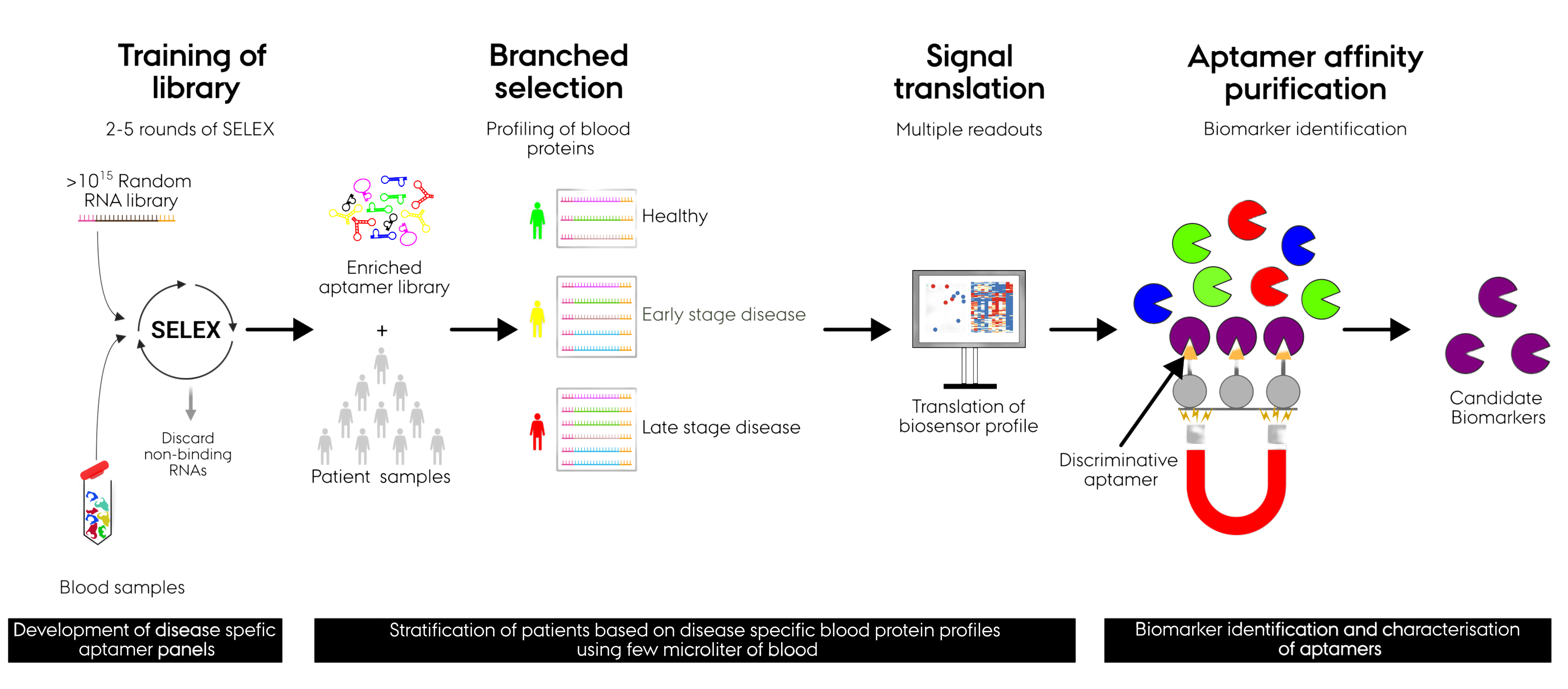
The AptaShape Method
Proteomics has proven to provide the most robust biomarkers in comparative studies but is complex, low-throughput and costly. Quantitative measurements of specific proteins commonly involve immunological methods, utilizing antibodies to detect protein epitopes in various assays.
Several screening tools for detecting biomarkers in biofluids have been pursued. While genetic approaches, such as circulating tumor DNA (ctDNA) can have predictive value, they often lack sensitivity for early cancer stages. Other approaches, such as circulating microRNA (transcriptomics), tumor cells (CTCs), metabolites and extracellular vesicles have also been reported as biomarkers but failed to reach general clinical use. Current proteomics array platforms based on pre-selected panels are biased towards the chosen probes and thereby “blind” to other proteins.
Our APTASHAPE method addresses these shortcomings by including a “training” step where a multitude of disease-specific protein biomarkers are identified prior to screening of patient groups. The general applicability and reproducibility of APTASHAPE have been validated across several disease indications using medium-sized independent cohorts (60-200 individuals).
APTASHAPE has been shown to predict healthy versus cancerous samples and distinguish early-stage from late-stage bladder cancer with 91% and 83% accuracy, respectively. Moreover, a defined panel of 25 aptamers from the bladder cancer aptamer library demonstrated a similar capacity to stratify healthy controls and cancer patients.
Proteomics has proven to provide the most robust biomarkers in comparative studies but is complex, low-throughput and costly. Quantitative measurements of specific proteins commonly involve immunological methods, utilizing antibodies to detect protein epitopes in various assays.
Several screening tools for detecting biomarkers in biofluids have been pursued. While genetic approaches, such as circulating tumor DNA (ctDNA) can have predictive value, they often lack sensitivity for early cancer stages. Other approaches, such as circulating microRNA (transcriptomics), tumor cells (CTCs), metabolites and extracellular vesicles have also been reported as biomarkers but failed to reach general clinical use. Current proteomics array platforms based on pre-selected panels are biased towards the chosen probes and thereby “blind” to other proteins.
Our APTASHAPE method addresses these shortcomings by including a “training” step where a multitude of disease-specific protein biomarkers are identified prior to screening of patient groups. The general applicability and reproducibility of APTASHAPE have been validated across several disease indications using medium-sized independent cohorts (60-200 individuals).
APTASHAPE has been shown to predict healthy versus cancerous samples and distinguish early-stage from late-stage bladder cancer with 91% and 83% accuracy, respectively. Moreover, a defined panel of 25 aptamers from the bladder cancer aptamer library demonstrated a similar capacity to stratify healthy controls and cancer patients.
